Abstract
Coral bleaching is the result of the algal symbiont being expelled from the gastrodermal tissue layer of coral polyps due to increased production of toxic reactive oxygen species (ROS) during photo oxidative stress events. Recent research has shown that certain beneficial bacteria can be applied as a probiotic supplement to reduce the effects of heat stress and mitigate the bleaching process. Exaiptasia diaphana (aiptasia), a common model organism used for coral research, was used in this experiment due to conservation protections placed on the endangered coral species. However, little is known about the microbiome of aiptasia which limits the potential for using aiptasia to research probiotics. This study isolated bacterial species most likely found in close association with the algal symbiont and one isolate was selected for further study based on morphology. The isolate was identified as a strain of Tritonibacter mobilis in NCBI using the 16S rRNA gene. To further confirm identity and enable future photo-oxidative stress assays, the Superoxide Dismutase (SOD) gene was located using degenerate primers designed from conserved and variable regions identified in an alignment of SOD sequences of closely related taxa. The SOD gene located in this strain of T. mobilis was identified as belonging to the Mn/Fe SOD gene family. By identifying culturable microbiome members that are closely associated with the algal symbiont and who have the potential to neutralize ROS, future research can delve more deeply into the mechanisms and functionality of coral probiotics during heat stress events.
Watch a recorded version of the presentation given at One USF Undergraduate Research Conference.
Introduction
Scleractinian coral are key ecosystem engineers (Jones et al., 1994) whose structural complexity provides habitat, shelter, and food for over 25% of all marine life (Graham and Nash 2013, Fisher et al. 2015, Fisheries 2023). This valuable ecosystem is critically endangered worldwide due to many stressors caused by climate change and especially rising ocean temperatures which have led to increasingly frequent bleaching events and subsequent coral mortality (Hutchins, 2025; National Oceanic and Atmospheric Administration, 2024). The scale and rate at which rising temperatures can trigger mass bleaching events makes this among the greatest threats to coral (Dörr et al., 2023; Palacio-Castro et al., 2022). Although there are many known causes of coral bleaching, the primary catalyst is acute heat stress and subsequent photooxidative stress which triggers termination of the symbiotic state between coral and its algal symbiont, either by expulsion, in-situ degradation, host cell detachment or apoptosis (Bieri et al., 2016; Gates et al., 1992). When the algal symbiont is gone, the skeleton is visible through the transparent coral tissue giving it a bleached white appearance (Voolstra, 2020). Given that the photoautotrophic Symbiodiniaceae provides over 90% of coral nutrients in an otherwise oligotrophic environment (Johnson, 2011), this often leads to mortality from starvation, disease, or parasites.
Photooxidative stress is the cellular damage caused when temperatures rise causing the PSII and the thylakoid membrane to break down (Deore et al., 2024) resulting in a buildup of toxic Reactive Oxygen Species (ROS) (Downs et al., 2002; Nielsen et al., 2018). Examples of ROS include superoxide (O₂•⁻), hydrogen peroxide (H₂O₂), and the hydroxyl radical (•OH), all of which form due to the incomplete reduction of oxygen (Toledano & Huang, 2016). As oxidative cellular damage accumulates, the rate of ROS production can increase by orders of magnitude and the holobiont’s antioxidant systems can be overwhelmed (Downs et al., 2002). The coral host will terminate the symbiosis in order to limit the damage caused by toxic ROS formation during a heat stress event resulting in coral bleaching. After a heat stress event the coral host must spend energy and resources on cellular and tissue repair. If the coral has lost its symbiont, there will be very little, if any, energy to spare rendering the coral highly susceptible to disease and parasites (Santoro et al., 2021).
Coral Probiotics
Coral probiotics is a relatively new field of research grounded in the Coral Probiotic Hypothesis that coral have evolved the ability to rapidly change their microbiome in response to environmental conditions in order to select for the most advantageous microbial partners (Voolstra & Ziegler, 2020). These advantageous microbial partners are commonly referred to as Beneficial Microorganisms for Coral (BMCs) and are defined as a collection of microorganisms that “contribute to coral health by aiding in nutrient uptake and growth, reducing the effects of stress and pollution or other toxins, having anti-pathogenic properties, and aiding in early life stage development” (Peixoto et al., 2017, 2021). Coral probiotics research uses this knowledge that many coral species can change their microbiome to design microbial interventions to mitigate stress and support coral resiliency.
There are many cited examples of success using probiotics to mitigate the effects of heat stress. One particularly compelling example was conducted in 2021, when Santoro et al. demonstrated the effect of BMCs during a simulated bleaching event in which every single coral genet in the placebo group either bleached bone white or died while the treatment group only paled or bleached but did not die and recovered significantly faster than the placebo group (Santoro et al., 2021). Another study found that BMC treatments given two hours prior to a heat stress event can mitigate that stress and prevent mortality (de Breuyn et al., 2025; Rosado et al., 2019). There have also been successful preliminary field tests to confirm probiotic effectiveness and a lack of off-target effects (Delgadillo-Ordoñez et al., 2024).
Our understanding of coral microbiome complexity by species, region, season, and habitat type has grown significantly with the development of metagenomics (Huggett & Apprill, 2019; Littman et al., 2010; Meron et al., 2011; Shaver et al., 2017; Ziegler et al., 2017). This understanding is critically important, as it gives a much more complete picture of the holobiont and all its members than a culture-based approach can, as this rarely captures more than a small fraction of the microbiome (Stewart, 2012; Sweet et al., 2021). However, metagenomics cannot replace the value of a culture-based approach to understanding bacterial metabolics where changes to environmental conditions can have pleiotropic effects on the holobiont that cannot be understood or evaluated using metagenomics alone (Sweet et al., 2021). Additionally, bacteria must actually be cultured in order to use them as a probiotic. In many cases, the limitations to culturing bacteria are due to a lack of understanding about the specific needs of a given bacterial species whether nutritional, environmental, or due to some as-yet-unknown inter- or intraspecies interactions (Stewart, 2012; Sweet et al., 2021).
The complexity of the coral microbiome requires a great deal more study. However, given the highly protected status of coral, many researchers use Exaiptasia diaphana, a common coral model organism. Currently, there is only limited research into probiotic consortia of Exaiptasia diaphana (hereafter referred to as aiptasia or by the specific clonal strain). Therefore, this study uses clonal strain CC7 aiptasia to isolate and identify BMC candidates that are native to this organism and that demonstrate photoprotective properties. The clonal strain CC7 is male and most likely originated in the Florida Keys (originally obtained from Carolina Biological Supply) (Dungan et al., 2020; Sunagawa et al., 2009) and contains Symbiodinium linucheae (ITS2 type A4) (LaJeunesse et al., 2018; Tortorelli et al., 2020).
Microbiome Compartments
It is well established over the past few decades that specific consortia of bacteria inhabit the various microhabitats or ‘compartments’ of coral anatomy; the surface mucus layer, the epidermal layer, the gastrodermal layer, the endolithic skeleton (Agostini et al., 2012; Glasl et al., 2016; Rosenberg et al., 2007), and the membrane-enclosed compartment or symbiosome where the symbiodinium are found (Gardner et al., 2023). Bacterial cultures from specific compartments can provide insight into biological or metabolic processes as well as interactions between the coral, its symbiont, and the environment (Armitage & Jones, 2019; Hughes et al., 2022).
Although all microbiome members play an important role in overall coral health, the consortia associated with the Symbiodinium have a greater impact on the holobiont as this is where corals derive nutrients and energy for reef building and where ROS are formed during heat stress events (Garrido et al., 2021; Maire et al., 2023; Muller-Parker et al., 2015; Sydnor et al., 2023).
Symbiodiniaceae are found in the symbiosome, a membranous vacuole located in the gastrodermal cells of the host (Garrido et al., 2021; Maire et al., 2023) (fig. 1). The Symbiodiniaceae phycosphere is the area within the symbiosome surrounding the Symbiodinium and is rich in nutrients excreted by the algal cell and with metabolites produced by a variety of bacteria also found there (Garrido et al., 2021).
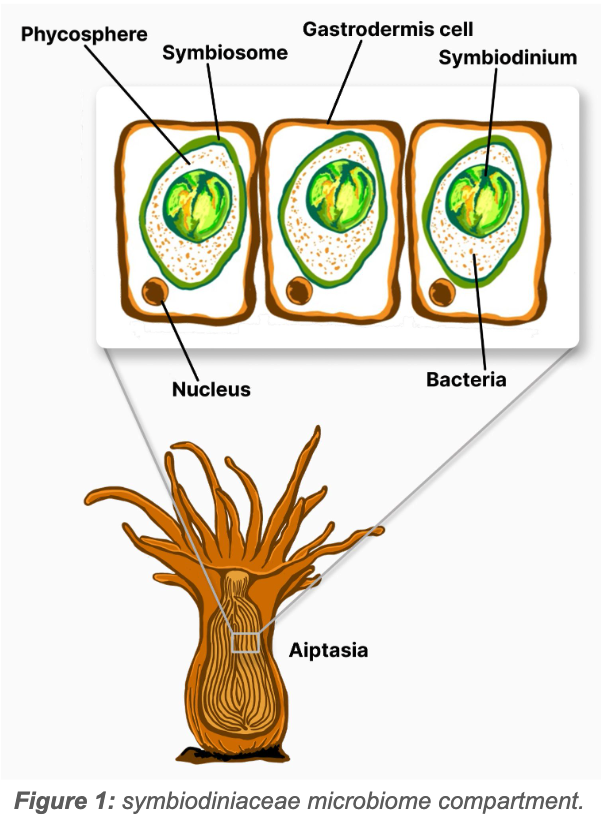
Research has shown that the bacterial assemblage most closely associated with the symbiodiniaceae phycosphere have greater Free Radical Scavenging (FRS) and nutrient cycling capacities that are closely linked to the coral-Symbiodinium symbiosis (Gardner et al., 2023; Garrido et al., 2021; Maire et al., 2021, 2023; Muller-Parker et al., 2015).
Photoprotective Capacity of the Microbiome
Although the current scale, frequency, and rate of ROS production is a modern problem for corals, the existence of ROS is not a new problem. Approximately 2.4 bya, as earth’s atmosphere became oxygenated, the formation of ROS was something the earliest life forms had to cope with (Case, 2017; Sheng et al., 2014). Several cellular response (anti-oxidant) systems have evolved to deal with toxic ROS production. These anti-oxidant systems can be found in the symbiont cells, the host cells, and the bacterial microbiome members (Fig. 2).
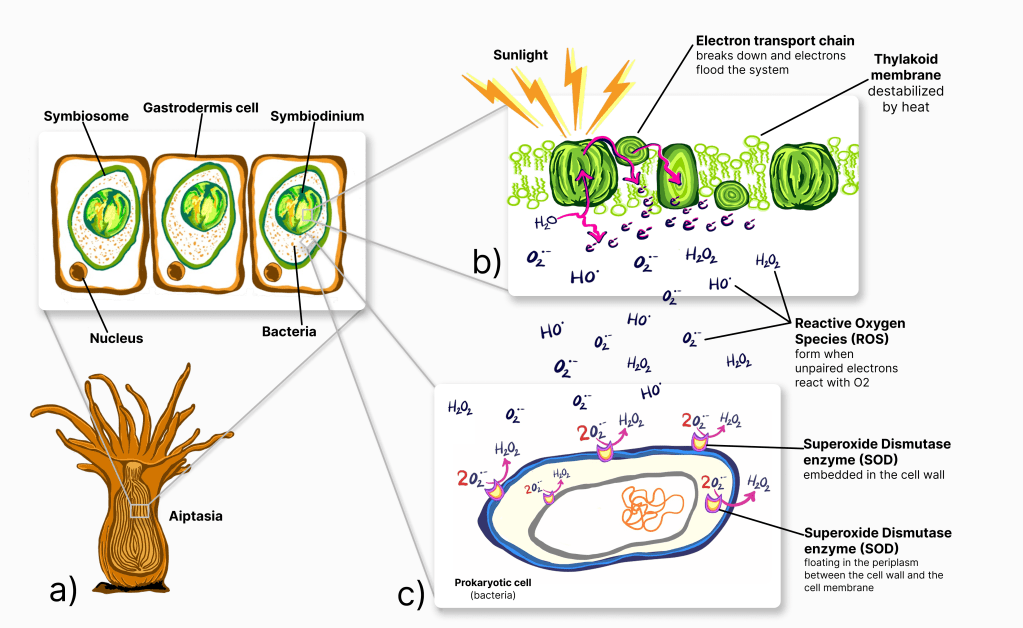
Figure 2: a) Coral polyps receive over 90% of their nutrition from their algal symbiont, Symbiodinium via photosynthesis. b) During a heat event, the thylakoid membrane, where photosynthesis takes place can destabilize, disrupting the photosynthesis electron transport chain (ETC). When the ETC breaks, unpaired electrons flood the area and react with nearby O₂ molecules to form highly toxic Reactive Oxygen Species (ROS). c) The Superoxide Dismutase Enzyme converts highly toxic superoxide ions (O₂•–) into less toxic hydrogen peroxide. Bacteria in the symbiosome that have extra-cellular or periplasmic SOD enzymes can mitigate the photo oxidative stress caused by excessive heat.
The most common anti-oxidant systems are peroxiredoxin (Prx), glutathione peroxidase (Gpx), superoxide dismutase (SOD), and Catalase (Bao et al., 2023). The peroxiredoxins (Prx) are actually a group of highly conserved thiol-based peroxidases that are ubiquitous among prokaryotes and have an extremely fast reaction rate when ROS levels are low (Toledano & Huang, 2016). Prx can act as a signalling molecule by temporarily deactivating to allow ROS to accumulate and thus activate additional anti-oxidant defense mechanisms (Rhee et al., 2012). While Prx is ubiquitous in prokaryotes, Glutathione peroxidase (Gpx) is more common in mammals and should be classified as “Trx-dependant, Gpx-like” though little is known about this highly diverse enzyme (Bao et al., 2023).
Unlike Gpx, superoxide dismutase (SOD) is very well studied in prokaryotes and converts the highly toxic superoxide (O2∙⁻) into O2 and H2O2 (Lynch & Kuramitsu, 2000). There are two forms of the SOD enzyme that are differentiated by the atom of metal used as a cofactor to stabilize the active site; one form uses either Mn or Fe and the other form uses Cu and Zn (Lynch & Kuramitsu, 2000). The performance of these SOD types are not equal; MnSOD has shown much higher protective capacity than FeSOD, but FeSOD has been shown to have a much steadier, more predictable level of activity (Lynch & Kuramitsu, 2000). The second type, Cu-ZnSOD, is less well understood in prokaryotes as it was once thought to be only found in Eukaryotes but has since been found in the periplasm and is thought to be active only during the stationary growth phase (Lynch & Kuramitsu, 2000). SOD activity can be highly variable in prokaryotes as some species may have one, none, or all SOD types present (Lynch & Kuramitsu, 2000).
Finally, catalase is a substrate specific enzyme that can rapidly convert H2O2 into H2O and O2, and is thought to be one of the earliest anti-oxidant enzymes to evolve (Iwase et al., 2013; Sheng et al., 2014). Catalase and SOD form a natural partnership in which SOD converts superoxides into O2 and H2O2 and Catalase converts H2O2 into H2O and O2. Given the highly conserved, ubiquitous nature of Prx and the less well-studied or understood status of Gpx in prokaryotes, this study focused on the Superoxide Dismutase and catalase activity of the microbiome.
This study attempted to isolate and culture bacterial species found within the symbiosome using isopycnic centrifugation with a discontinuous Percoll gradient followed by filtering and serial dilutions. The SOD gene for the cultured isolate was located using primers designed from the conserved regions of the SOD gene in related species and the SOD gene type was identified using key identifying characteristics in the amino acid sequence.
By isolating and identifying beneficial microbiome members of aiptasia CC7 that are most closely associated with the algal symbiont, future research can delve more deeply into the mechanisms and functionality of coral probiotics during heat stress events using these model organisms.
Materials and Methods
Aiptasia clonal strain CC7 was kept in a 3.8 L tank with a small bubbler filter in Instant Ocean Artificial Seawater (ASW) at a salinity level of 35ppt and kept at temperatures between 25℃ – 27℃. Light was provided at approximately 70 µmol·m-2·s-1 for 12 hours each day. Aiptasia were fed brine shrimp once each week and tanks were cleaned with a complete water change after feeding. The bubbler portion of the filter was gently scrubbed to remove visible algae and detritus and the sponge filter was gently squeezed in clean ASW to remove large bits of detritus only. Aiptasia were not fed 1 week prior to experimental protocol to avoid contamination from brine shrimp.
Tissue sample collection
A tissue sample was collected from three aiptasia polyps whose oral disks were at least 1cm in diameter. The polyps were rinsed in sterile artificial seawater (SASW) and then macerated using a centrifuge tube pestle. The macerated tissue was further homogenized using a 19 gauge needle for approximately ten passes and then a 20 gauge needle for approximately ten passes until the sample was a homogenous slurry.
Cell separation
In order to separate the host tissue from the Symbiodinium, a discontinuous gradient of Percoll solution was used based on protocols described by Hill et al., 2024; MilliporeSigma, n.d.; Peng et al., 2010; Schultz et al., 2022; Ziegler et al., 2017. A Stock Isotonic Percoll (SIP) solution consisting of nine parts Percoll and one part 5.5 M solution of NaCl in Deionized water was created by first adding 0.322 g of NaCl and 1 mL of DI water to a conical tube. Once most of the NaCl was dissolved, the Percoll solution was added and the contents were mixed by gently inverting the conical tube several times to allow the remaining NaCl to dissolve evenly in the solution. For this experiment, no Osmometer was available so the osmolarity was confirmed via calculations. See supplemental materials for calculation details.
The following concentrations of SIP were created by mixing SIP and SASW to create 100% SIP, 80% SIP, 50% SIP, and 25% SIP solutions. A discontinuous gradient was built by layering 2 mL of each SIP concentration in the following order: 100%, 80%, 50%, and 25%. The homogenized aiptasia tissue suspended in SASW was layered on top of the gradient (fig. 3a) and then centrifuged at 5,000×g for 15 minutes. After centrifugation, the host tissue formed a mostly clear, colorless, and highly viscous layer at the top with a dark, clumpy layer of Symbiodinium lower in the gradient (fig. 3b). In order to verify the gradient remained present after centrifugation, a small amount of food coloring was added to separate samples of the same gradient concentrations. Fig. 3c shows that the gradient remained stable with minimal mixing.
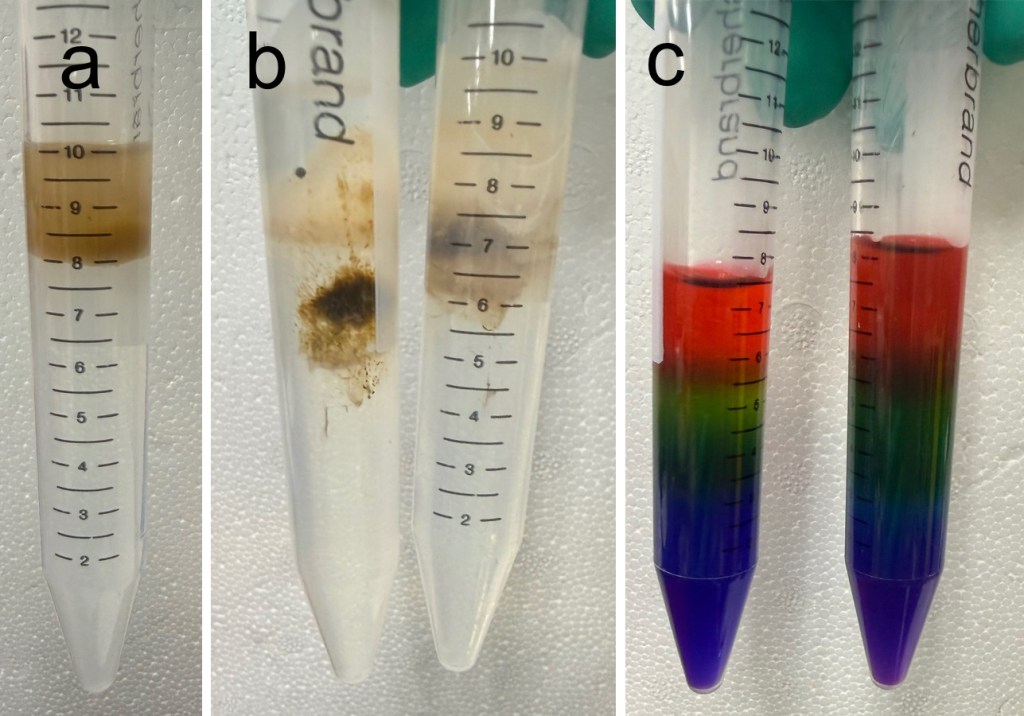
Figure 3: a) centrifuge tube with SIP gradient layers and tissue slurry on top pre-centrifugation. b) centrifuge tubes with separated cell types after centrifugation. c) verification of gradient stability after centrifugation.
The Symbiodinium were further cleaned and separated based on protocols outlined by Hill et al., 2024; and Maire et al., 2021, 2023. The fraction containing the Symbiodinium cells was filtered through a 30 µm, 20 µm, and a 15 µm filter to remove large debris and any remaining host cells (fig. 4 a-d). The final filter was 5 µm and small enough to catch the majority of the Symbiodinium cells which were then resuspended in 1 mL of SASW. The resuspended cells were then diluted up to 10-4 and plated on 2x dilute Zobell’s marine broth amended with 1 g/L of glucose. All plates were then placed in an incubator at 26℃ for 2-3 days.
One isolate was selected based on colony morphology (fig. 4e) and plated using the 4-quadrant streak plate technique to ensure a pure culture was isolated.
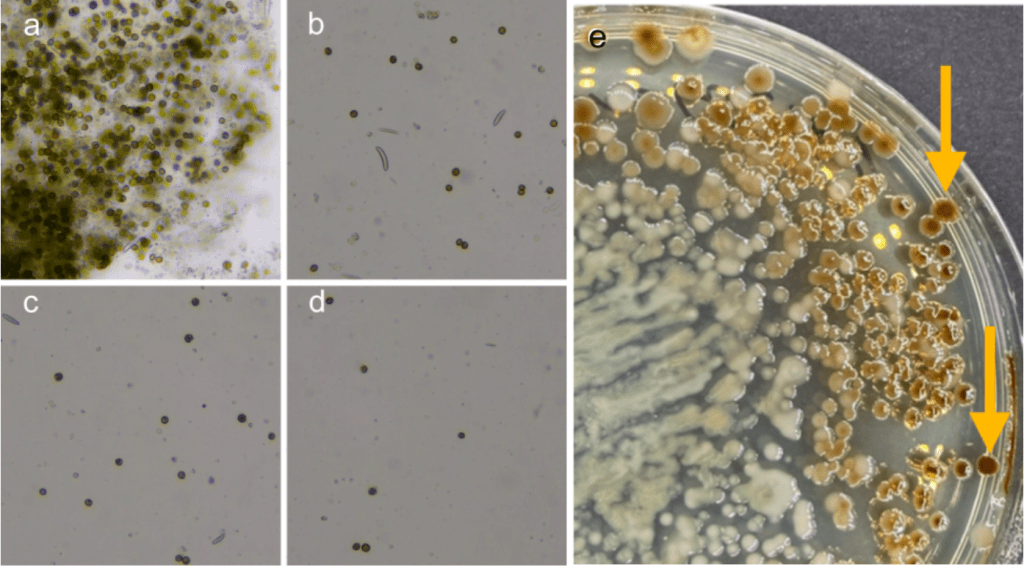
Figure 4: a) symbiodinium cells immediately after separation. Filtered cells after being passed through; b) a 30 µm filter, c) a 20 µm filter, d) a 15 µm filter. e) Initial plate results from plated Symbiodinium cells. Arrows indicate examples of colony morphology for the selected isolate.
Species Identification with the 16S rRNA gene
DNA was extracted using Zymo Research’s Quick-DNA Fungal/Bacterial Microprep kit. DNA was amplified using Thermo Scientific DreamTaq Green PCR Master mix and sequenced by Eurofins Genomics using Sanger sequencing.
The sequenced results were run through the NCBI database using the BLAST tool to identify the isolate as Tritonibacter mobilis. However, because the NCBI search results listed six different closely related species and strain matches with ≥ 97% identity, there was some degree of uncertainty as to the species and strain-level identification. In order to resolve this, additional steps were taken to locate the Superoxide Dismutase gene.
Locating and identifying the Superoxide Dismutase gene
In order to identify the Superoxide Dismutase (SOD) gene, a phylogenetic tree was generated in NCBI from the top 13 matches for the 16S gene and a more closely related subset was identified (yellow highlighted box in fig. 5).
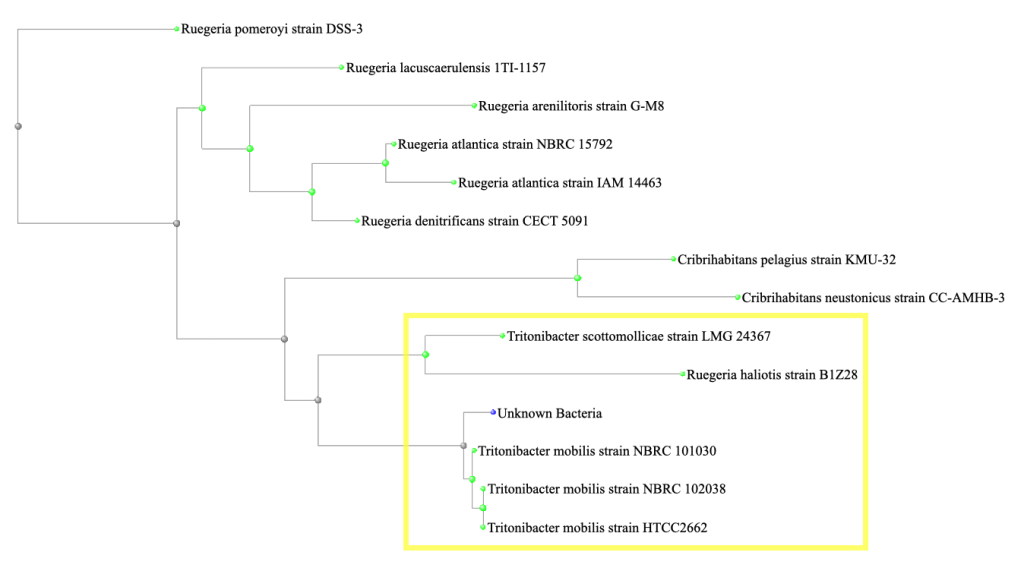
Figure 5: A phylogenetic tree generated in NCBI from the top 13 matches for the 16S gene. A more closely related subset is identified within the yellow highlighted box.
An alignment of the SOD gene sequences from Ruegeria haliotis, Tritonibacter scottomollicae, and three different strains of Tritonibacter mobilis was created using Clustal Omega. The forward primer (GCGACGTTTTCCCAGTTCAC) was designed using the conserved regions and a degenerate reverse primer (TGGCTTTTGAACTTCCYGAYC) was designed to accommodate shorter conserved regions.
The SOD gene was amplified via PCR (3 minutes at 95℃, then 35 cycles of 95℃ for one minute, 55℃ for one minute, and 72℃ for two minutes, followed by 72℃ for 5 minutes) and sequenced by Eurofins Genomics using Sanger sequencing.
The SOD gene nucleotide sequence was converted to its amino acid sequence using the ExPASy translate tool and 3-D protein models were generated using ColabFold, AlphaFold2 notebook and visualized in the RCSB Protein Data Bank.
Results
The Superoxide Dismutase gene matched seven different strains of Tritonibacter mobilis (Epibacterium mobile and Ruegeria mobilis are former names for T. mobilis) with ≥ 99% identity. All of the SOD strain matches were different from the strains matched by the 16S gene. Figure six shows the top four matches for the SOD gene with the nucleotide differences noted as red letters below the query sequence.
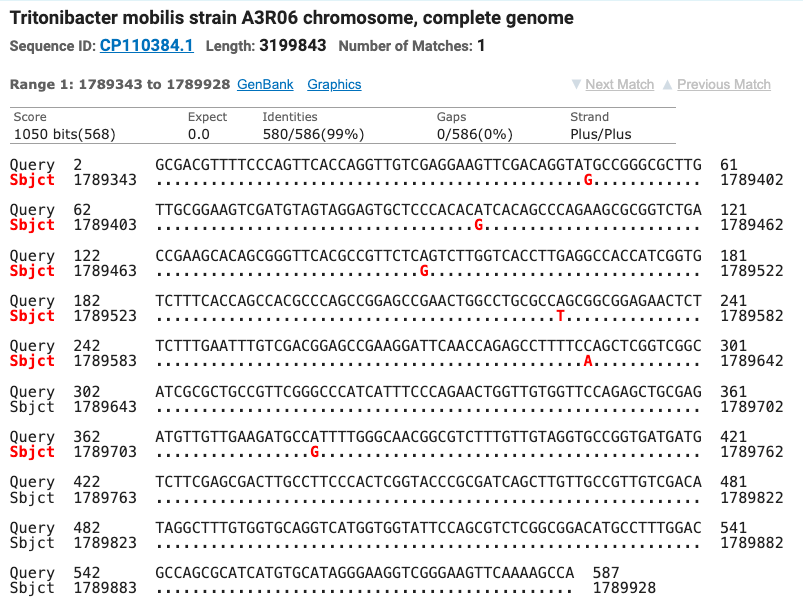
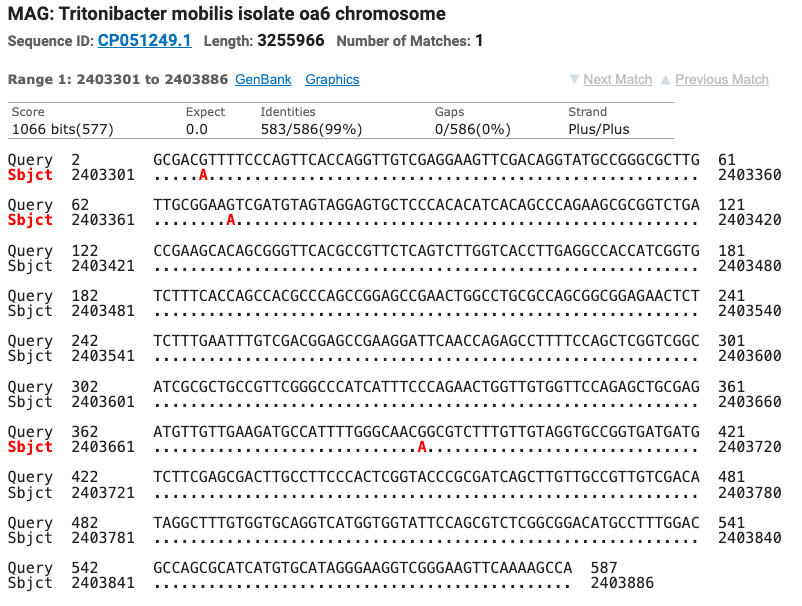
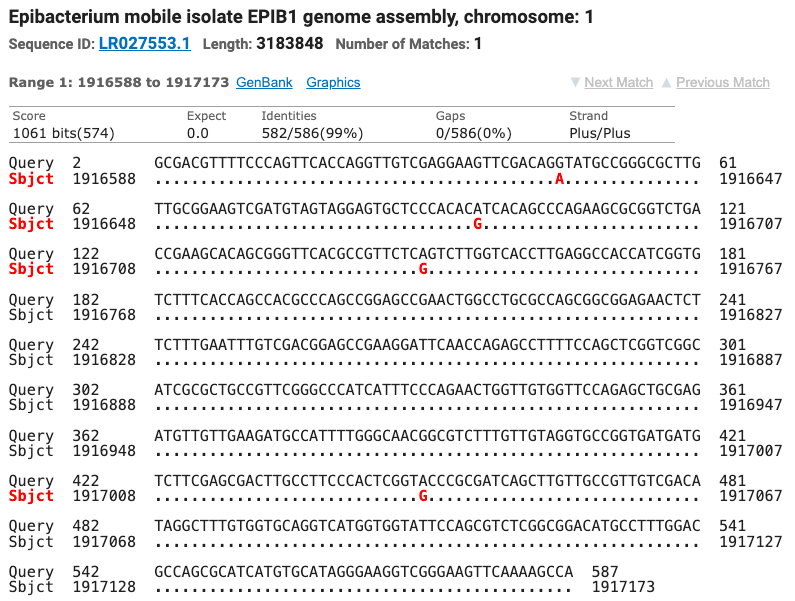
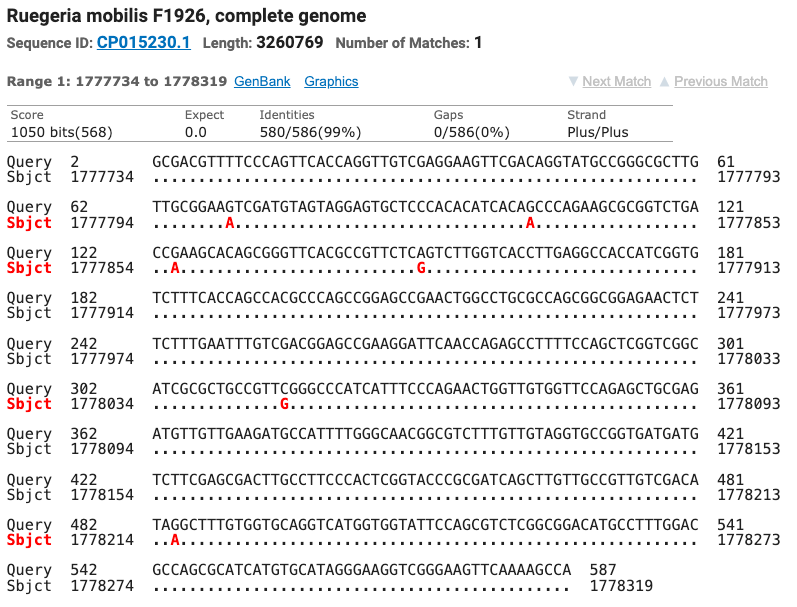

Figure 6: The top four matches for the isolate’s SOD gene. Red letters show the nucleotide differences between the isolate and the matched species.
The sequenced results for the forward primer and the reverse primer were checked against the electropherogram and aligned to create the most complete SOD gene sequence possible with the available sequence results. The full SOD sequence is shown in figure 7 above the longest open reading frame found by the ExPASy translate tool.

Figure 7: Top; full nucleotide sequence for the isolate’s SOD gene, Bottom; Longest open reading frame found by the ExPASy translate tool.
The specific type of SOD gene found in this isolate was identified based on two key features of the amino acid sequence. First, the underlined sequence with the yellow highlight box in fig. 8a is highly conserved in the Manganese/Iron SOD gene family. Second, the relative spacing of the four specific amino acids highlighted in yellow also support the Mn/Fe SOD gene family identity. When the protein is folded – those yellow highlighted amino acids will form the active binding site for the metal ion that stabilizes and catalyzes the SOD enzyme.
Fig. 8b shows the structure of the folded protein and fig. 8c shows that same protein with the active binding site formed by those key amino acids highlighted in red.
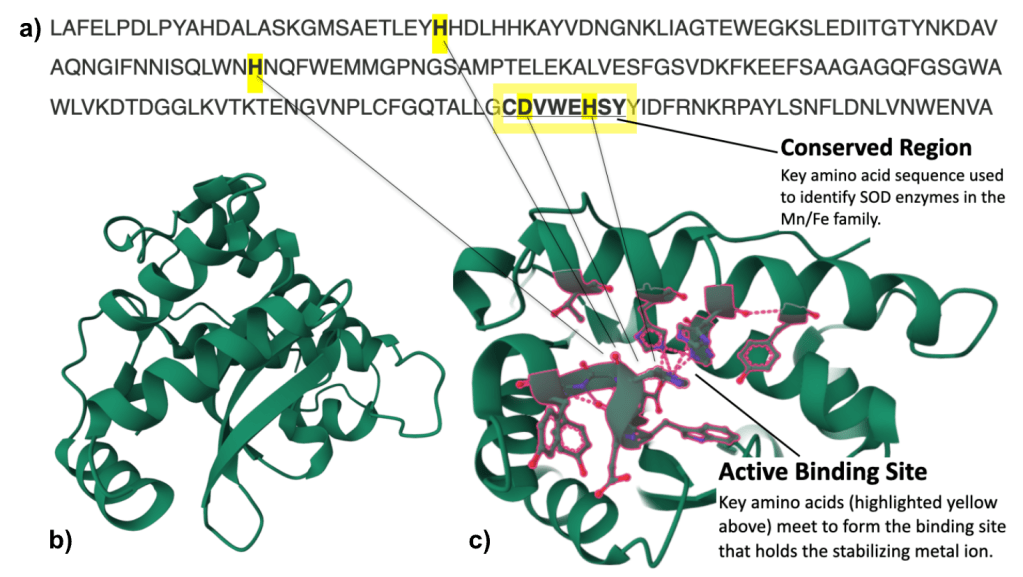
Figure 8: a) SOD amino acid sequence converted from nucleotide sequence using the ExPASy translate tool. b) SOD protein modeled in 3-D. c) SOD protein with active binding site amino acids highlighted in red. Both 3-D protein models were generated using ColabFold, AlphaFold2 notebook and visualized in RCSB Protein Data Bank.
Discussion
A single isolate was selected for this study and taken from initial isolation to species identification, location of the SOD gene, and verification of the Mn/Fe SOD gene family. Although no SOD enzyme activity measurements were successfully taken, this study demonstrates a replicable process for culturing bacterial isolates most likely to be in close association with the Symbiodinium, identifying the isolates, and locating the SOD gene.
Ultimately, the purpose of this research is to investigate ways to slow the coral bleaching process and ultimately lower coral mortality rates during and after a bleaching event by identifying bacteria that are able to up regulate SOD enzyme production when it detects signs of photo-oxidative stress and then use these bacterial species as part of a probiotic mix administered to its host prior to heat stress.
In order to do this, the next phase of this research would involve measuring baseline SOD enzyme activity and SOD transcription levels in bacteria isolated from aiptasia. Then exposing bacteria to simulated photo-oxidative stress and measuring SOD enzyme activity and SOD transcription levels again. Any bacteria that demonstrate the ability to up regulate SOD enzyme production would be selected to be part of a probiotic mix for future heat stress testing experiments. Figure 9 illustrates these steps.
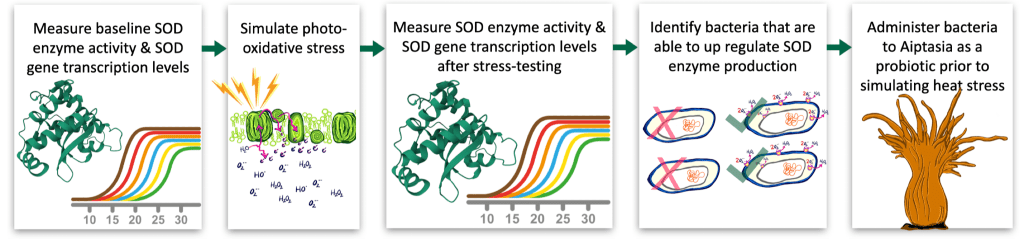
Figure 9: Next steps from left to right: Measure baseline SOD enzyme activity and gene transcription levels in bacteria isolated from aiptasia, expose isolates to simulated photo-oxidative stress, re-measure SOD activity and transcriptional response, select strains that demonstrate significant up-regulation of SOD, Incorporate selected strains into a candidate probiotic mix for future heat stress experiments.
By identifying culturable members of the microbiome that are closely associated with algal symbionts and exhibit the potential to neutralize reactive oxygen species, this work could provide a foundation for future studies to more deeply investigate the mechanisms and functional roles of coral probiotics during heat stress events.
Acknowledgements
I would like to express sincere gratitude to Dr. Cory Krediet (Eckerd College) for providing aiptasia specimens, and to Dr. Mya Breitbart (USFSTP College of Marine Science) for essential laboratory materials. I also thank Dr. Ana Hoare (USFSTP Chemistry Department) for her chemistry support. Additional thanks go to Lauren Winegarden, Cassandra Schneider, and Samantha Greenfield for their assistance during extended lab sessions. Finally, I am grateful to the University of South Florida Department of Integrative Biology and Professor Howard (Lab Manager) for supporting the MicroBioRes course that made this research possible, as well as the lab preparation staff for their help with media, cultures, and overall laboratory logistics.
Citations
Agostini, S., Suzuki, Y., Higuchi, T., Casareto, B. E., Yoshinaga, K., Nakano, Y., & Fujimura, H. (2012). Biological and chemical characteristics of the coral gastric cavity. Coral Reefs, 31(1), 147–156. https://doi.org/10.1007/s00338-011-0831-6
Armitage, D. W., & Jones, S. E. (2019). How sample heterogeneity can obscure the signal of microbial interactions. The ISME Journal, 13(11), 2639–2646. https://doi.org/10.1038/s41396-019-0463-3
Bao, Y.-J., Zhou, Q., Yu, X., Yu, X., & Castellino, F. J. (2023). Analysis and Characterization of Glutathione Peroxidases in an Environmental Microbiome and Isolated Bacterial Microorganisms. Journal of Microbiology and Biotechnology, 33(3), 299–309. https://doi.org/10.4014/jmb.2209.09006
Bieri, T., Onishi, M., Xiang, T., Grossman, A. R., & Pringle, J. R. (2016). Relative Contributions of Various Cellular Mechanisms to Loss of Algae during Cnidarian Bleaching. PLoS ONE, 11(4), e0152693. https://doi.org/10.1371/journal.pone.0152693
Case, A. J. (2017). On the Origin of Superoxide Dismutase: An Evolutionary Perspective of Superoxide-Mediated Redox Signaling. Antioxidants, 6(4), 82. https://doi.org/10.3390/antiox6040082
de Breuyn, M., Ostendarp, M., El-Khaled, Y. C., Garcias-Bonet, N., Carvalho, S., Wild, C., & Peixoto, R. S. (2025). Probiotics prevent mortality of thermal-sensitive corals exposed to short-term heat stress. ISME Communications, 5(1), ycaf039. https://doi.org/10.1093/ismeco/ycaf039
Delgadillo-Ordoñez, N., Garcias-Bonet, N., Raimundo, I., García, F. C., Villela, H., Osman, E. O., Santoro, E. P., Curdia, J., Rosado, J. G. D., Cardoso, P., Alsaggaf, A., Barno, A., Antony, C. P., Bocanegra, C., Berumen, M. L., Voolstra, C. R., Benzoni, F., Carvalho, S., & Peixoto, R. S. (2024). Probiotics reshape the coral microbiome in situ without detectable off-target effects in the surrounding environment. Communications Biology, 7(1), 434. https://doi.org/10.1038/s42003-024-06135-3
Deore, P., Tsang Min Ching, S. J., Nitschke, M. R., Rudd, D., Brumley, D. R., Hinde, E., Blackall, L. L., & van Oppen, M. J. H. (2024). Unique photosynthetic strategies employed by closely related Breviolum minutum strains under rapid short-term cumulative heat stress. Journal of Experimental Botany, 75(13), 4005–4023. https://doi.org/10.1093/jxb/erae170
Dörr, M., Denger, J., Maier, C. S., Kirsch, J. V., Manns, H., & Voolstra, C. R. (2023). Short-term heat stress assays resolve effects of host strain, repeat stress, and bacterial inoculation on Aiptasia thermal tolerance phenotypes. Coral Reefs, 42(6), 1271–1281. https://doi.org/10.1007/s00338-023-02427-y
Downs, C. A., Fauth, J. E., Halas, J. C., Dustan, P., Bemiss, J., & Woodley, C. M. (2002). Oxidative stress and seasonal coral bleaching. Free Radical Biology and Medicine, 33(4), 533–543. https://doi.org/10.1016/S0891-5849(02)00907-3
Dungan, A. M., Bulach, D., Lin, H., Oppen, M. J. H. van, & Blackall, L. L. (2020). Development of a free radical scavenging probiotic to mitigate coral bleaching (p. 2020.07.02.185645). bioRxiv. https://doi.org/10.1101/2020.07.02.185645
Fisher, R., O’Leary, R. A., Low-Choy, S., Mengersen, K., Knowlton, N., Brainard, R. E., & Caley, M. J. (2015). Species Richness on Coral Reefs and the Pursuit of Convergent Global Estimates. Current Biology, 25(4), 500–505. https://doi.org/10.1016/j.cub.2014.12.022
Gardner, S. G., Leggat, W., & Ainsworth, T. D. (2023). The microbiome of the endosymbiotic Symbiodiniaceae in corals exposed to thermal stress. Hydrobiologia, 850(17), 3685–3704. https://doi.org/10.1007/s10750-023-05221-7
Garrido, A. G., Machado, L. F., Zilberberg, C., & De Assis Leite, D. C. (2021). Insights into “Symbiodiniaceae phycosphere” in a coral holobiont. Symbiosis, 83, 25–39. https://doi.org/10.1007/s13199-020-00735-3
Gates, R., Baghdasarian, G., & Muscatine, L. (1992, June). Temperature Stress Causes Host Cell Detachment in Symbiotic Cnidarians: Implications for Coral Bleaching. https://doi.org/10.2307/1542252
Glasl, B., Herndl, G. J., & Frade, P. R. (2016). The microbiome of coral surface mucus has a key role in mediating holobiont health and survival upon disturbance. The ISME Journal, 10(9), 2280–2292. https://doi.org/10.1038/ismej.2016.9
Graham, N. A. J., & Nash, K. L. (2013). The importance of structural complexity in coral reef ecosystems. Coral Reefs, 32(2), 315–326. https://doi.org/10.1007/s00338-012-0984-y
Hill, L. J., Messias, C. S. M. de A., Vilela, C. L. S., Garritano, A. N., Villela, H. D. M., do Carmo, F. L., Thomas, T., & Peixoto, R. S. (2024). Bacteria associated with the in hospite Symbiodiniaceae’s phycosphere. iScience, 27(4), 109531. https://doi.org/10.1016/j.isci.2024.109531
Huggett, M. J., & Apprill, A. (2019). Coral microbiome database: Integration of sequences reveals high diversity and relatedness of coral‐associated microbes. Environmental Microbiology Reports, 11(3), 372–385. https://doi.org/10.1111/1758-2229.12686
Hughes, D. J., Raina, J.-B., Nielsen, D. A., Suggett, D. J., & Kühl, M. (2022). Disentangling compartment functions in sessile marine invertebrates. Trends in Ecology & Evolution, 37(9), 740–748. https://doi.org/10.1016/j.tree.2022.04.008
Hutchins, R. (2025, July 4). Drone footage reveals Australia’s “unprecedented” coral mortality. Oceanographic. https://oceanographicmagazine.com/news/drone-footage-reveals-australia-s-unprecedented-coral-mortality/
Iwase, T., Tajima, A., Sugimoto, S., Okuda, K., Hironaka, I., Kamata, Y., Takada, K., & Mizunoe, Y. (2013). A Simple Assay for Measuring Catalase Activity: A Visual Approach. Scientific Reports, 3, 3081. https://doi.org/10.1038/srep03081
Johnson, M. D. (2011). The acquisition of phototrophy: Adaptive strategies of hosting endosymbionts and organelles. Photosynthesis Research, 107(1), 117–132. https://doi.org/10.1007/s11120-010-9546-8
Jones, C. G., Lawton, J. H., & Shachak, M. (1994). Organisms as Ecosystem Engineers. Oikos, 69(3), 373. https://doi.org/10.2307/3545850
LaJeunesse, T. C., Parkinson, J. E., Gabrielson, P. W., Jeong, H. J., Reimer, J. D., Voolstra, C. R., & Santos, S. R. (2018). Systematic Revision of Symbiodiniaceae Highlights the Antiquity and Diversity of Coral Endosymbionts. Current Biology, 28(16), 2570-2580.e6. https://doi.org/10.1016/j.cub.2018.07.008
Littman, R. A., Bourne, D. G., & Willis, B. L. (2010). Responses of coral-associated bacterial communities to heat stress differ with Symbiodinium type on the same coral host. Molecular Ecology, 19(9), 1978–1990. https://doi.org/10.1111/j.1365-294X.2010.04620.x
Lynch, M., & Kuramitsu, H. (2000). Expression and role of superoxide dismutases (SOD) in pathogenic bacteria. Microbes and Infection, 2(10), 1245–1255. https://doi.org/10.1016/S1286-4579(00)01278-8
Maire, J., Girvan, S. K., Barkla, S. E., Perez-Gonzalez, A., Suggett, D. J., Blackall, L. L., & Van Oppen, M. J. H. (2021). Intracellular bacteria are common and taxonomically diverse in cultured and in hospite algal endosymbionts of coral reefs. The ISME Journal, 15(7), 2028–2042. https://doi.org/10.1038/s41396-021-00902-4
Maire, J., Tandon, K., Collingro, A., Van De Meene, A., Damjanovic, K., Gotze, C. R., Stephenson, S., Philip, G. K., Horn, M., Cantin, N. E., Blackall, L. L., & Van Oppen, M. J. H. (2023). Colocalization and potential interactions of Endozoicomonas and chlamydiae in microbial aggregates of the coral Pocillopora acuta. Science Advances, 9(20), eadg0773. https://doi.org/10.1126/sciadv.adg0773
Meron, D., Atias, E., Iasur Kruh, L., Elifantz, H., Minz, D., Fine, M., & Banin, E. (2011). The impact of reduced pH on the microbial community of the coral Acropora eurystoma. The ISME Journal, 5(1), 51–60. https://doi.org/10.1038/ismej.2010.102
MilliporeSigma. (n.d.). How to Make and Use Percoll Gradients. Retrieved September 3, 2025, from https://www.sigmaaldrich.com/US/en/technical-documents/protocol/cell-culture-and-cell-culture-analysis/mammalian-cell-culture/how-to-make-and-use-gradients-of-percoll
Muller-Parker, G., D’Elia, C. F., & Cook, C. B. (2015). Interactions Between Corals and Their Symbiotic Algae. In C. Birkeland (Ed.), Coral Reefs in the Anthropocene (pp. 99–116). Springer Netherlands. https://doi.org/10.1007/978-94-017-7249-5_5
National Oceanic and Atmospheric Administration. (2024, April 15). NOAA confirms 4th global coral bleaching event |. https://www.noaa.gov/news-release/noaa-confirms-4th-global-coral-bleaching-event
Nielsen, D. A., Petrou, K., & Gates, R. D. (2018). Coral bleaching from a single cell perspective. The ISME Journal, 12(6), 1558–1567. https://doi.org/10.1038/s41396-018-0080-6
NOAA. (2023, January). How are Fisheries and Coral Reefs Connected? (National). NOAA. http://www.fisheries.noaa.gov/feature-story/how-are-fisheries-and-coral-reefs-connected
Palacio-Castro, A. M., Rosales, S. M., Dennison, C. E., & Baker, A. C. (2022). Microbiome signatures in Acropora cervicornis are associated with genotypic resistance to elevated nutrients and heat stress. Coral Reefs, 41(5), 1389–1403. https://doi.org/10.1007/s00338-022-02289-w
Peixoto, R. S., Rosado, P. M., Leite, D. C. de A., Rosado, A. S., & Bourne, D. G. (2017). Beneficial Microorganisms for Corals (BMC): Proposed Mechanisms for Coral Health and Resilience. Frontiers in Microbiology, 8. https://doi.org/10.3389/fmicb.2017.00341
Peixoto, R. S., Sweet, M., Villela, H. D. M., Cardoso, P., Thomas, T., Voolstra, C. R., Høj, L., & Bourne, D. G. (2021). Coral Probiotics: Premise, Promise, Prospects. Annual Review of Animal Biosciences, 9(1), 265–288. https://doi.org/10.1146/annurev-animal-090120-115444
Peng, S., Wang, Y., Wang, L., Chen, W. U., Lu, C., Fang, L., & Chen, C. (2010). Proteomic analysis of symbiosome membranes in Cnidaria–dinoflagellate endosymbiosis. PROTEOMICS, 10(5), 1002–1016. https://doi.org/10.1002/pmic.200900595
Rhee, S. G., Woo, H. A., Kil, I. S., & Bae, S. H. (2012). Peroxiredoxin Functions as a Peroxidase and a Regulator and Sensor of Local Peroxides*. Journal of Biological Chemistry, 287(7), 4403–4410. https://doi.org/10.1074/jbc.R111.283432
Rosado, P. M., Leite, D. C. A., Duarte, G. A. S., Chaloub, R. M., Jospin, G., Nunes Da Rocha, U., Saraiva, J. P., Dini-Andreote, F., Eisen, J. A., Bourne, D. G., & Peixoto, R. S. (2019). Marine probiotics: Increasing coral resistance to bleaching through microbiome manipulation. The ISME Journal, 13(4), 921–936. https://doi.org/10.1038/s41396-018-0323-6
Rosenberg, E., Koren, O., Reshef, L., Efrony, R., & Zilber-Rosenberg, I. (2007). The role of microorganisms in coral health, disease and evolution. Nature Reviews Microbiology, 5(5), 355–362. https://doi.org/10.1038/nrmicro1635
Santoro, E. P., Borges, R. M., Espinoza, J. L., Freire, M., Messias, C. S. M. A., Villela, H. D. M., Pereira, L. M., Vilela, C. L. S., Rosado, J. G., Cardoso, P. M., Rosado, P. M., Assis, J. M., Duarte, G. A. S., Perna, G., Rosado, A. S., Macrae, A., Dupont, C. L., Nelson, K. E., Sweet, M. J., … Peixoto, R. S. (2021). Coral microbiome manipulation elicits metabolic and genetic restructuring to mitigate heat stress and evade mortality. Science Advances, 7(33), eabg3088. https://doi.org/10.1126/sciadv.abg3088
Schultz, J., Modolon, F., Rosado, A. S., Voolstra, C. R., Sweet, M., & Peixoto, R. S. (2022). Methods and Strategies to Uncover Coral-Associated Microbial Dark Matter. mSystems, 7(4), e00367-22. https://doi.org/10.1128/msystems.00367-22
Shaver, E. C., Shantz, A. A., McMinds, R., Burkepile, D. E., Vega Thurber, R. L., & Silliman, B. R. (2017). Effects of predation and nutrient enrichment on the success and microbiome of a foundational coral. Ecology, 98(3), 830–839. https://doi.org/10.1002/ecy.1709
Sheng, Y., Abreu, I. A., Cabelli, D. E., Maroney, M. J., Miller, A.-F., Teixeira, M., & Valentine, J. S. (2014). Superoxide Dismutases and Superoxide Reductases. Chemical Reviews, 114(7), 3854–3918. https://doi.org/10.1021/cr4005296
Stewart, E. J. (2012). Growing Unculturable Bacteria. Journal of Bacteriology, 194(16), 4151–4160. https://doi.org/10.1128/jb.00345-12
Sunagawa, S., Wilson, E. C., Thaler, M., Smith, M. L., Caruso, C., Pringle, J. R., Weis, V. M., Medina, M., & Schwarz, J. A. (2009). Generation and analysis of transcriptomic resources for a model system on the rise: The sea anemone Aiptasia pallida and its dinoflagellate endosymbiont. BMC Genomics, 10(1), 258. https://doi.org/10.1186/1471-2164-10-258
Sweet, M., Villela, H., Keller-Costa, T., Costa, R., Romano, S., Bourne, D. G., Cárdenas, A., Huggett, M. J., Kerwin, A. H., Kuek, F., Medina, M., Meyer, J. L., Müller, M., Pollock, F. J., Rappé, M. S., Sere, M., Sharp, K. H., Voolstra, C. R., Zaccardi, N., … Peixoto, R. (2021). Insights into the Cultured Bacterial Fraction of Corals. mSystems, 6(3), e01249-20. https://doi.org/10.1128/mSystems.01249-20
Sydnor, J. R., Lopez, J., Wolfe, G. V., Ott, L., & Tran, C. (2023). Changes in the microbiome of the sea anemone Exaiptasia diaphana during bleaching from short-term thermal elevation. Frontiers in Marine Science, 10, 1130964. https://doi.org/10.3389/fmars.2023.1130964
Toledano, M. B., & Huang, B. (2016). Microbial 2-Cys Peroxiredoxins: Insights into Their Complex Physiological Roles. Molecules and Cells, 39(1), 31–39. https://doi.org/10.14348/molcells.2016.2326
Tortorelli, G., Belderok, R., Davy, S. K., McFadden, G. I., & van Oppen, M. J. H. (2020). Host Genotypic Effect on Algal Symbiosis Establishment in the Coral Model, the Anemone Exaiptasia diaphana, From the Great Barrier Reef. Frontiers in Marine Science, 6. https://doi.org/10.3389/fmars.2019.00833
Voolstra, C. R. (2020). Coral Bleaching: A Colorful Struggle for Survival. Current Biology, 30(13), R768–R770. https://doi.org/10.1016/j.cub.2020.05.008
Voolstra, C. R., & Ziegler, M. (2020). Adapting with Microbial Help: Microbiome Flexibility Facilitates Rapid Responses to Environmental Change. BioEssays, 42(7), 2000004. https://doi.org/10.1002/bies.202000004
Ziegler, M., Seneca, F. O., Yum, L. K., Palumbi, S. R., & Voolstra, C. R. (2017). Bacterial community dynamics are linked to patterns of coral heat tolerance. Nature Communications, 8(1), 14213. https://doi.org/10.1038/ncomms14213


